Why Google Cloud
Why Google Cloud

Scale with infrastructure built for performance
Citadel Securities accelerates quantitative research 2-4x faster at 30% less cost with Google Cloud TPUs.

Turn fragmented customer service into seamless journeys
The Home Depot drives higher conversion rates from inspiration to installation with Gemini Enterprise for Customer Experience.

Empower your crew with a world-class AI assistant
Virgin Voyages uses Gemini Enterprise on Google Distributed Cloud Edge to deliver instant, personalized service regardless of satellite connectivity.

Innovate smartly with an integrated AI stack
Unilever connects agents end-to-end so procurement teams can make faster, smarter sourcing decisions on a secure foundation.
Product Category
- AI and Machine Learning
- API Management
- Compute
- Containers
- Data Analytics
- Databases
- Developer Tools
- Google Workspace
- Networking
- Operations
- Security and Identity
- Serverless Computing
- Storage
- Other
Industry
- Automotive
- Education
- Financial & Insurance Services
- Gaming
- Government & Public Sector
- Healthcare
- Life Sciences
- Manufacturing
- Media & Entertainment
- Retail & Consumer Goods
- Startup
- Technology
- Telecommunications
- Transportation
- Other
Region
- Asia Pacific & Japan
- Europe, Middle East & Africa
- Latin America
- U.S. & Canada
- AI and Machine Learning
- API Management
- Compute
- Containers
- Data Analytics
- Databases
- Developer Tools
- Google Workspace
- Networking
- Operations
- Security and Identity
- Serverless Computing
- Storage
- Other
- Automotive
- Education
- Financial & Insurance Services
- Gaming
- Government & Public Sector
- Healthcare
- Life Sciences
- Manufacturing
- Media & Entertainment
- Retail & Consumer Goods
- Startup
- Technology
- Telecommunications
- Transportation
- Other
- Asia Pacific & Japan
- Europe, Middle East & Africa
- Latin America
- U.S. & Canada
Product Category
Industry
Region
- Relevance
- Newest
- Title A-Z
 "Etsy creates personalized experiences for its 90 million shoppers by using Vertex AI, BigQuery, Dataflow, and Gemini models to better understand inventory, customer intent, and individual buyers."
"Etsy creates personalized experiences for its 90 million shoppers by using Vertex AI, BigQuery, Dataflow, and Gemini models to better understand inventory, customer intent, and individual buyers." Wayfair embraces Google's Gemini models and Google AI, powering new experiences that make it easier for anyone, anywhere to create their feeling of home.
Wayfair embraces Google's Gemini models and Google AI, powering new experiences that make it easier for anyone, anywhere to create their feeling of home. UCR partnered with Google Public Sector to deploy Gemini Enterprise, automating routine workflows to create a more intelligent, agent-powered campus that supports students and empowers faculty.
UCR partnered with Google Public Sector to deploy Gemini Enterprise, automating routine workflows to create a more intelligent, agent-powered campus that supports students and empowers faculty. CME Group uses Gemini Code Assist to accelerate developer tasks and enable its technical workforce to focus on innovation.
CME Group uses Gemini Code Assist to accelerate developer tasks and enable its technical workforce to focus on innovation. "A concept we're embracing as an organization is having digital co-workers, enabled by Gemini Enterprise. It makes sense as the next step in incorporating AI as an integral part of how we work."
"A concept we're embracing as an organization is having digital co-workers, enabled by Gemini Enterprise. It makes sense as the next step in incorporating AI as an integral part of how we work." PayPal adopted Looker's Model Context Protocol (MCP) Server to provide secure, conversational analytics to its workforce, overcoming complex security challenges by automating credential encryption. By grounding AI interactions in Looker's governed data models, PayPal ensures accuracy and compliance while enabling 5,000 employees to query petabytes of data using natural language.
PayPal adopted Looker's Model Context Protocol (MCP) Server to provide secure, conversational analytics to its workforce, overcoming complex security challenges by automating credential encryption. By grounding AI interactions in Looker's governed data models, PayPal ensures accuracy and compliance while enabling 5,000 employees to query petabytes of data using natural language. The Indiana Secretary of State (SOS) partnered with Google Public Sector to transform 20M+ records into living intelligence, launch bilingual AI agents, and deploy an award-winning LMS for 50k+ statewide notaries. Using Gemini models, Indiana SOS is revolutionizing resident services and support.
The Indiana Secretary of State (SOS) partnered with Google Public Sector to transform 20M+ records into living intelligence, launch bilingual AI agents, and deploy an award-winning LMS for 50k+ statewide notaries. Using Gemini models, Indiana SOS is revolutionizing resident services and support. Thrive is using BigQuery and Gemini with the goal of cutting data integration time by 50% to help restaurant managers focus more on customer and team experiences.
Thrive is using BigQuery and Gemini with the goal of cutting data integration time by 50% to help restaurant managers focus more on customer and team experiences. HeyGen uses Gemini to power its Video Agent, reducing video creation from days to minutes and delivering professional quality videos.
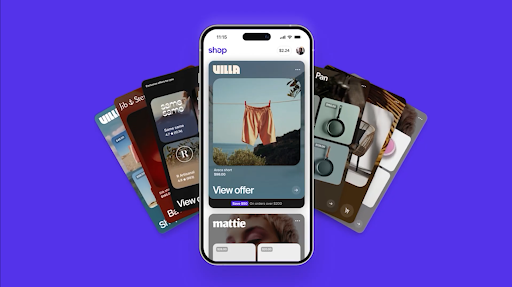
HeyGen uses Gemini to power its Video Agent, reducing video creation from days to minutes and delivering professional quality videos. Shopify uses Google Cloud to process $14.6B in Black Friday weekend sales without compromising speed and reliability for 81M buyers.
Shopify uses Google Cloud to process $14.6B in Black Friday weekend sales without compromising speed and reliability for 81M buyers. With Google AI, Leaseo is replacing fragmented property management systems with a single, cloud-based AI platform that streamlines and automates key processes.
With Google AI, Leaseo is replacing fragmented property management systems with a single, cloud-based AI platform that streamlines and automates key processes. Boys Town, a 501(c)(3) nonprofit helping at-risk youth and families, launched Reality Coach, a simulation coaching and training platform, on Google Cloud to scale evidence-based behavioral training, empowering educators and improving classroom experiences and outcomes for students.
Boys Town, a 501(c)(3) nonprofit helping at-risk youth and families, launched Reality Coach, a simulation coaching and training platform, on Google Cloud to scale evidence-based behavioral training, empowering educators and improving classroom experiences and outcomes for students. Cartesia built the world's fastest, ultra-realistic voice AI on Google Cloud, delivering sub-90ms latency and 99.99% uptime to 50K+ companies—transforming how the world speaks, listens, and does business across millions of conversations every day.
Cartesia built the world's fastest, ultra-realistic voice AI on Google Cloud, delivering sub-90ms latency and 99.99% uptime to 50K+ companies—transforming how the world speaks, listens, and does business across millions of conversations every day.
Learn what industry analysts are saying about Google Cloud
Explore analyst reportsNetwork with other professionals
Join the Google Cloud community
Ask about Google Cloud
- Clear conversation
- replyDiscover solutions for my industry
- replyHow do companies use Gemini Enterprise?
- replyTell me 5 innovative things customers do with Google Cloud