Getting to know the Google Cloud Healthcare API: part 3
Chris von See
Staff Technical Solutions Consultant
The Google Cloud Healthcare & Life Sciences team recently introduced a set of application programming interfaces (APIs) and datastores that can be used to enable healthcare data analysis and machine learning applications, data-level integration of healthcare systems and secure storage and retrieval of various types of electronic patient healthcare information (ePHI). The first article in this series provided a brief overview of the Cloud Healthcare API and its possible use cases, and the second article explained some of the concepts implemented in the API and illustrated basic approaches to using the API. In this final segment we’ll do a deeper dive into how you can use the API to address challenges you might have in your own organization.
About our use case
A key use case that is becoming increasingly relevant in many healthcare organizations is generating, storing and surfacing machine learning predictions on FHIR, HL7v2 and DICOM data. Because FHIR and DICOM play a significant role in both the path to ML predictions and as sources for valuable data, we’ll also show how these data modalities affect the process of implementing machine learning on healthcare data.
Architecture foundations
Before diving into an explanation of the core architecture that makes the magic happen, let’s discuss some foundational work that needs to occur before loading sensitive data into Google Cloud Platform (GCP).
Preparing to work with PHI in Google Cloud
The Google Cloud documentation provides guidance regarding considerations and best practices for complying with regulatory requirements that govern how to handle personal healthcare information (PHI). While we won’t reproduce all of that information here, it’s important to highlight some of the more important tasks you should perform before implementing any PHI processing in GCP:
- In the United States, ensure that you have a signed HIPAA Business Associates Agreement (BAA), and that you have explicitly disabled non-compliant products and services in your GCP projects. Google Cloud provides open-source deployment scripts that can assist with correct project setup, and our documentation includes a list of HIPAA-compliant GCP services for US deployments.
- Create Identity and Access Management (IAM) service accounts and keys for processing PHI data, and grant those accounts the least amount of privilege they need to perform their work. The Cloud Healthcare API has a comprehensive set of available IAM roles and permissions that you can use, and you can combine these with roles and permissions for other GCP services to create the right combination for your needs.
- If you are using Google Cloud Storage to hold PHI data—either as a data ingestion point or as part of a data lake strategy—ensure that the data is accessible only to people and systems explicitly authorized to access it. When using “gsutil” to upload and/or download data, be aware of the security and privacy considerations that can help keep your data safe.
- Determine if your data requires encryption above and beyond what Google Cloud itself provides. Google provides a cloud-managed encryption key (CMEK) service that can be useful in certain cases, such as encrypting data in Google Cloud Storage—check the product documentation to determine if the GCP data services you’re planning to use support CMEK and if so, how best to implement it.
- Activate Google Cloud audit logging for both administrative and data events, and export this data periodically so that it can be reviewed for possible security breaches. Google BigQuery can be very useful for analyzing this kind of log data. As always, avoid logging sensitive PHI data by applications that consume this data in order to avoid inadvertent exposure to unauthorized personnel.
- To further control access to sensitive data you may want to consider limiting access to systems that process PHI data by implementing a virtual private clouds (VPCs). VPCs can be subdivided into one or more public or private networks, and you can implement a number of strategies to prevent access from the public internet.
- When it’s necessary to exchange sensitive information with non-GCP systems or applications, virtual private network (VPN), mutual TLS (mTLS) and the Apigee API management system are great options. One or more of these techniques can be used to meet your specific access control requirements.
Setting up authentication for Google Cloud
All of the modality-specific Cloud Healthcare APIs require authentication via Google Cloud Identity and Access Management (IAM). To set up authentication for the Cloud Healthcare API, create an IAM service account and assign Cloud Healthcare API roles that are appropriate for the type of access required. For service accounts that will be used for administrative activities, for example, appropriate roles include those that enable dataset and store creation (typically, the "Healthcare Dataset Administrator" and "Healthcare <modality> Store Administrator" roles); other roles will be appropriate for other use cases (the “Healthcare HL7V2 Ingest” role for sending data to the HL7v2 store, for example). Be sure to grant only the lowest level of permissions required for the given use case.
Cloud Healthcare API authentication requires the use of OAuth 2 access tokens, which can be generated in a number of different ways. The easiest way to do this from an interactive (command-line) session is to use the “gcloud application-default print-access-token” function. Download the service account JSON key and store it in a secure location, then create an environment variable called GOOGLE_APPLICATION_CREDENTIALS with a value specifying the full path to the downloaded key. This environment variable will be used by the “gcloud” command to create or retrieve OAuth 2 access tokens and to authenticate subsequent calls to the Cloud Healthcare API.
Note that sometimes, applications that issue Cloud Healthcare API requests cannot use “gcloud” to retrieve Oauth access tokens. In that case, you can use the same service account key to generate tokens using the open source “oauth2l” utility, or use Apigee in combination with the open-source Cloud Healthcare API token generator.
Generating, storing and surfacing machine learning predictions on FHIR, HL7v2 or DICOM data
Leveraging machine learning to streamline healthcare workflows or gain insights into data is an emerging use case that holds great promise for addressing some of the critical challenges faced by the healthcare industry.
Baseline architecture
The diagram below represents a baseline architecture for FHIR, HL7v2 and DICOM data integration. It shows three integration patterns:
- Use of an integration engine such as Cloverleaf or NextGen Connect (formerly Mirth Connect). This can be appropriate for both FHIR and HL7v2 data.
- Direct use of the Cloud Healthcare API MLLP adapter to forward data to the Cloud Healthcare HL7v2 API
- Ingestion of DICOM imaging data from a PACS system or vendor-neutral archive.
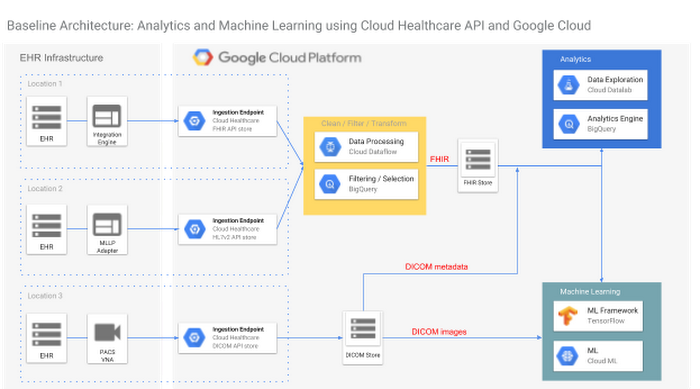

Each type of data goes through three phases: ingestion, cleansing / filtering / transformation, and processing. The details of each step, however, differ somewhat depending on the data modality.
These patterns are by no means the only ways to ingest these types of data—the best methods for your particular situation may differ from this example. For example, it’s possible to extract data directly from the EHR’s datastore, transfer it to Google Cloud Storage in a bulk format such as CSV, and then ingest it into Cloud Dataflow or BigQuery in the cleaning / filtering / transformation stage. This can be particularly valuable when real-time updates aren’t needed.
Ingesting HL7v2 data
Generally, when ingesting HL7v2 data you will use an integration engine such as NextGen Connect or Cloverleaf to send requests to the MLLP adapter or to make requests to the Cloud Healthcare HL7v2 API directly. Google provides an open-source MLLP adapter that accepts inbound connections and forwards the HL7v2 data sent on the connection to the Cloud Healthcare HL7v2 API, where it is securely stored in an HL7v2 store. This approach might be appropriate if you do not plan to modify the data before ingestion. While the Cloud Healthcare HL7v2 API is a cloud-only service, the adapter itself can be hosted either in your own data center or in the cloud.
If you do plan to generate or edit HL7v2 messages yourself using an integration engine, you can invoke the Cloud Healthcare HL7v2 API directly using your integration engine’s REST API invocation feature. Similar to the MLLP adapter, the Cloud Healthcare HL7v2 API accepts and stores the complete HL7v2 message, and it can interpret some elements of the structure and content such as the message header, patient IDs, etc., so that you can filter messages you read from the store.
Ingesting FHIR data
FHIR data can be imported into Google Cloud using the Cloud Healthcare FHIR API. This API supports the FHIR DSTU2, STU3, and R4 standard, making it easy to bring both individual FHIR entities or bundles of related FHIR data into GCP. The Cloud Healthcare FHIR API can be invoked either directly by applications (as might be the case with a bulk load operation) or via an integration engine. Once ingested, the data is stored in a secure FHIR store, where it can be made available for analysis or for application access.
Ingesting DICOM data
DICOM studies are imported into Google Cloud via the Cloud Healthcare DICOMweb API, which natively complies with the DICOMweb standard (an open-source DIMSE adapter that implements STOW-RS operations is also available). Because DICOM studies are composed of both image data and metadata, making effective use of this data often requires separate processing paths for the metadata and image data. The Cloud Healthcare DICOMweb API provides features that allow you to export the metadata into an analytics system such as BigQuery, as well as image-related facilities that support de-identification of data and bulk export for deep machine learning analysis.
Cleansing, filtering and transformation
In order to get the best possible results when analyzing healthcare data or obtaining predictions from machine learning models, you should transform your data into a standardized schema. This is particularly true for HL7v2 data, which is difficult to use for analysis and prediction because of the time-based, transaction-oriented nature of the messages and the large amount of variability in message content. As a standard with a large amount of structural consistency across entities, FHIR is a good candidate for a target schema; Google has focused on making both BigQuery and machine learning tools that can perform analysis, inferences and predictions on FHIR data.
To streamline the process of converting HL7v2 data, the Cloud Healthcare HL7v2 API parses the HL7 “pipe-and-hat” format into a more easily consumable JSON structure.
Data processing
Google Cloud provides several different ways to apply machine learning to healthcare data. BigQuery’s powerful built-in machine learning capability (which is in beta at this writing) enables execution of models for linear regression, binary logistic regression or multiclass logistic regression from within the SQL language, vastly simplifying use for data analysts performing these types of predictions. For more complex predictions or insights, organizations can develop their own machine-learning models using systems such as TensorFlow, and can execute these models using TensorFlow Serving or Google Cloud ML Engine. Machine learning is particularly interesting when used with radiological images; predictions on these images provide an opportunity to streamline radiological workflows by categorizing and prioritizing studies that require review by a trained imaging specialist.
FHIR data (whether ingested natively in this format or transformed from HL7v2) can be exported from the Cloud Healthcare API FHIR store and imported into BigQuery for analysis using BigQuery’s SQL commands. When data is exported from a Cloud Healthcare FHIR store into BigQuery, a series of tables are created, each of which corresponds to a different FHIR entity type: Patient, Observation, Encounter, and so on. Using SQL, these tables can be examined by data scientists to gain deep insight into the nature of the data. Further, BigQuery SQL allows data analysts to join the data across tables in order to answer many different types of questions about trends in patient care, medication usage, readmission rates, and so on. Combining FHIR data with metadata extracted from DICOM studies using the Cloud Healthcare DICOM API can provide even more insight.
Implementing analytics and machine learning prediction
You can use the Cloud Healthcare API to import data and forward it automatically through each of the steps we described above. To do this, you will need to enable the Cloud Healthcare API in your projects.
We’ll use the following conventions for any sample Cloud Healthcare API requests:
“PROJECT_ID” refers to the ID of your Google Cloud project.
“LOCATION” refers to the name of the Google Cloud region in which you place your data.
“DATASET_ID” is the name of the Cloud Healthcare API dataset that contains your modality-specific store and data.
“STORE_TYPE” is the type of your modality-specific store. Valid values are “hl7V2Stores”, “fhirStores” and “dicomStores” for HL7v2, FHIR and DICOM data storage, respectively.
“STORE_ID” is the name of your modality-specific store.
Determine the source of your data
As a first step you should determine the source of your data—the EHR itself, an interface engine, a PACS system, an outside service, etc. If you plan to use the MLLP adapter for HL7v2 data then you should install and configure it in an appropriate location—see the MLLP adapter Github project for more details on how to do this. You may also want to set up a VPN to enable more secure transmission of healthcare data to Google Cloud; the process for setting up a VPN is outside the scope of this document but information is available in the Google Cloud VPN documentation.
Sending data to a Cloud Healthcare API store
Part 2 of this blog series talked in detail about the concept of Cloud Healthcare API datasets and stores, and gave some examples illustrating how to create them. In this section we’ll assume that you’ve created a dataset and modality-specific store, and that you’re now ready to insert data into that store to make it available for analysis, inferences or predictions.
Regardless of where your data originates, you will use a Cloud Healthcare API request to send data into the modality-specific datastore you created previously. The specific syntax of each request is modality-specific; requests to store DICOM data, for example, use a request syntax based on the DICOMweb standard, and requests to store FHIR data use request syntax based on the FHIR standard. You should consult the relevant standard for your modality, as well as the Cloud Healthcare API documentation, to determine the specific requests needed to ingest data into your modality-specific store, but generally all requests will follow a format similar to the one shown below:
where the “<modality-specific-healthcare-data>” value in the POST data contains the data you wish to ingest, and “<MODALITY_SPECIFIC_REQUEST>” is the modality-specific portion of the request. For example, a request to ingest a FHIR “Patient” entity into a FHIR store called “myStore” might look like this:
Similarly, a request to ingest an HL7v2 message might look like this:
where the “<base64-encoded-hl7v2-message>” value in the POST data contains a complete, Base64-encoded HL7v2 message.
Some modalities—DICOM, for example—also provide bulk “import” and “export” requests that allow for larger-scale ingestion of data in that modality. For details on how to use these bulk requests, consult the Cloud Healthcare API documentation.
Cleaning, filtering and transforming modality-specific data
If the modality-store has been configured with a Cloud Pub/Sub topic, applications subscribed to that topic will receive a notification of available data. This notification will contain a reference to the corresponding entity in the modality-specific store, and you can use that reference to read the entity for processing. You can handle the Pub/Sub notifications and read the relevant entities in an application running in Google Kubernetes Engine or Compute Engine, in a Cloud Functions service, or directly inside a Cloud Dataflow pipeline by using the Apache Beam “PubsubIO” adapters for Google Cloud (available for the Java, Python and Go programming languages).
If your data is in HL7v2 format, the process of converting to FHIR is specific to your particular HL7v2 usage and your desired FHIR content mapping. Currently, this conversion and mapping process is done using custom logic running in Cloud Dataflow or another GCP service. Once the data is transformed into its target FHIR schema you can store it in a Cloud Healthcare API FHIR store. This enables you to both use the data for analysis, inference and predictions, and also to serve both the data and the results of processing—stored in FHIR format—to applications.
Loading data into BigQuery and performing analysis
Cloud Healthcare FHIR API makes it very simple to move FHIR data into BigQuery for analysis. Using the “export” API request, you can specify a GCP project ID and BigQuery dataset, and the Cloud Healthcare FHIR API will create the appropriate BigQuery tables and schemas and start an operation to export the data into the selected dataset. Once that operation is complete the data can then be queried using BigQuery’s SQL functions.
To start an export operation, use an API request similar to the following:
where “<BQ_PROJECT_ID>” is the name of the GCP project containing the BigQuery instance to which you wish to export data, and “<BQ_DATASET_ID>” is the ID of the BigQuery dataset into which to export the data. This returns an operation request ID, which you can then query using the Cloud Healthcare API “operations” function:
where “<OPERATION_ID>” is the ID of the export operation as indicated in the “export” API response. When the export operation is complete, the response from the “operations” request will include a property “done”:true to indicate that the operation has completed.
Note that in order to export data to BigQuery the Cloud Healthcare API must have “job create” access to BigQuery. You can grant permission to create BigQuery jobs to the Cloud Healthcare API by looking for a service account in Cloud IAM with the name “Cloud Healthcare Service Agent” and then assigning the “BigQuery Job User” role to that account.
Invoking machine learning models and capturing the results
While there are no clinically-approved, pre-trained models for DICOM image analysis available from Google, experiences such as our work in detecting diabetic retinopathy or Stanford University’s work to identify skin cancer illustrate the promise of applying machine learning to imaging studies. Pre-screening DICOM studies with machine learning models can, for example, support and focus the work of radiologists by enabling them to concentrate on those studies that display the highest probability of disease or that exhibit characteristics that are of concern.
FHIR data stored in a Cloud Healthcare API FHIR store can be used by machine learning models to obtain inferences or predictions about the data. To do this, you can use Cloud Functions to invoke your model via either TensorFlow Serving or Cloud ML Engine. The results from the model can then be stored in FHIR format inside the same FHIR store as the source data, and identifiers such as the patient’s Medical Records Number (MRN) enable the model output to be matched to the corresponding patient. Google has provided a basic open-source example of the integration between Cloud Healthcare FHIR API and machine learning on Github.
Examples of FHIR entities which might be used to record results or drive actions based on inferences or predictions include:
RiskAssessment, for assessments of likely outcomes and the probability of each outcome.
Condition, possibly with a “provisional” condition verification status
ProcedureRequest, using a “proposal” request intent code to suggest procedures that might lead to a diagnosis without explicitly authorizing those procedures
This post on the Google AI Blog gives a brief overview of TensorFlow Serving, describing how it can be deployed and used to create inferences in production environments. TensorFlow Serving supports the use of a gRPC interface to enable applications to invoke the model, making it easy to integrate via Cloud Functions or other applications.
Similar to TensorFlow Serving, Cloud ML Engine provides an API that lets you get predictions out of your model. Cloud ML Engine’s documentation provides some specific guidance for obtaining predictions in online systems with low latency, which might be appropriate for situations where you are ingesting data in near-real-time via a Cloud Healthcare API FHIR store and the Cloud Pub/Sub topic. For batch processing—where you batch data in a Cloud Healthcare API FHIR store and process it in bulk—there is an alternative batch processing model that is well suited to this approach.
Conclusions and next steps
The Cloud Healthcare API and GCP give healthcare IT a powerful suite of tools to organize healthcare data and to make it accessible, secure and useful. Leveraging the Cloud Healthcare API to improve interoperability across care systems opens up opportunities for powerful new applications that take advantage of deep analytics, artificial intelligence, and massive computing and storage scale. If your organization can benefit from making information more accessible, we invite you to explore the Cloud Healthcare API and explore Google’s healthcare offerings to see how we can help.



