AI Platform Notebooks now supports R in beta

Karthik Ramachandran
Product Manager, Google Cloud
At Next ‘19, we announced the beta availability of AI Platform Notebooks, our managed service that offers an integrated environment to create JupyterLab instances that come pre-installed with the latest data science and machine learning frameworks. Today, we’re excited to introduce support for R on AI Platform Notebooks. You can now spin up a web-based development environment with JupyterLab, IRkernel, xgboost, ggplot2, caret, rpy2 and other key R libraries pre-installed.
The R language is a powerful tool for data science, and has been popular with data engineers, data scientists, and statisticians everywhere since its first release in 1992. It offers a sprawling collection of open source libraries that contain implementations of a huge variety of statistical techniques. For example, the Bioconductor library contains state of the art tools for analyzing genomic data. Likewise, with the forecast package you can carry out very sophisticated time series analysis using models like ARIMA, ARMA, AR, and exponential smoothing. Or, if you prefer building deep learning models, you could use TensorFlow for R.
Users of R can now leverage AI Platform Notebooks to create instances that can be accessed via the web or via SSH. This means you can install the libraries you care about; and you can easily scale your notebook instances up or down.
Getting started is easy
You can get started by navigating to the AI Platform and clicking on Notebooks. Then:
1. Click on “New Instance” and select R 3.5.3 (the first option).
2. Give your instance a name and hit “Create”.
In a few seconds your Notebook instance will show up in the list of instances available to you.
You can access the instance by clicking on “Open JupyterLab”.
This brings up the JupyterLab Launcher. From here you can do these three things:
1. Create a new Jupyter Notebook using IRKernel by clicking on the R button under Notebook.
2. Bring up an iPython style console for R by clicking on the R button under Console.
3. Open up a terminal by clicking on the terminal button under Other.
For fun, let’s create a new R notebook and visualize the infamous ‘Iris’ dataset, which consists of the measurements of the size of various parts of an Iris labeled by the particular species of Iris. It’s a good dataset for trying out simple clustering algorithms.
1. Create a new R notebook by clicking on the R button under Notebooks.
2. In the first cell, type in:
data(iris)head(iris)This will let you see the first 6 rows of the Iris data set.
3. Next, let’s plot Petal.Length against Sepal.Length:library(‘ggplot2’)ggplot(iris, aes(x = Petal.Length, y = Sepal.Length, colour = Species)) +
geom_point() +
ggtitle('Iris Species by Petal and Sepal Length')


Install additional R packages
As mentioned earlier, one of the reasons for R’s popularity is the sheer number of open source libraries available. One popular package hosting service is the Comprehensive R Archive Network (CRAN), with over 10,000 published libraries.
You can easily install any of these libraries from the R console. For example, if you wanted to install the widely popular igraph—a package for doing network analysis—you could do so by opening up the R console and running the install.packages command:
install.packages(“igraph”)
Scale up and down as you need
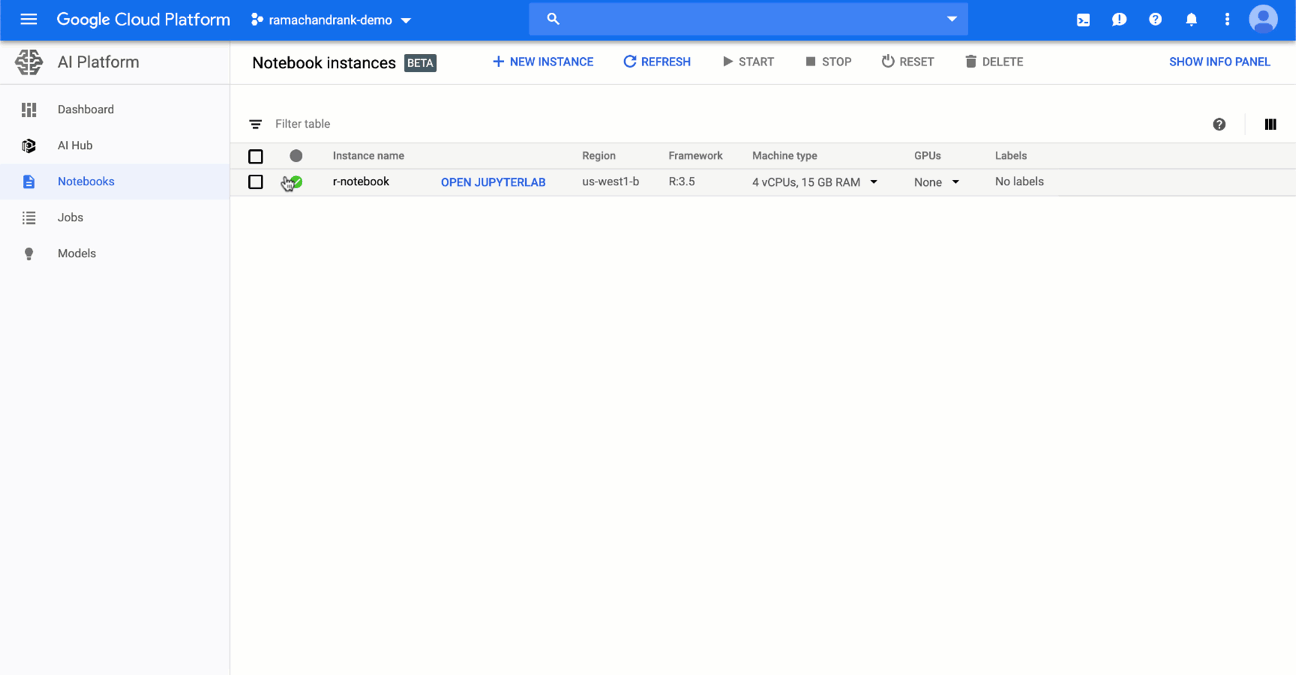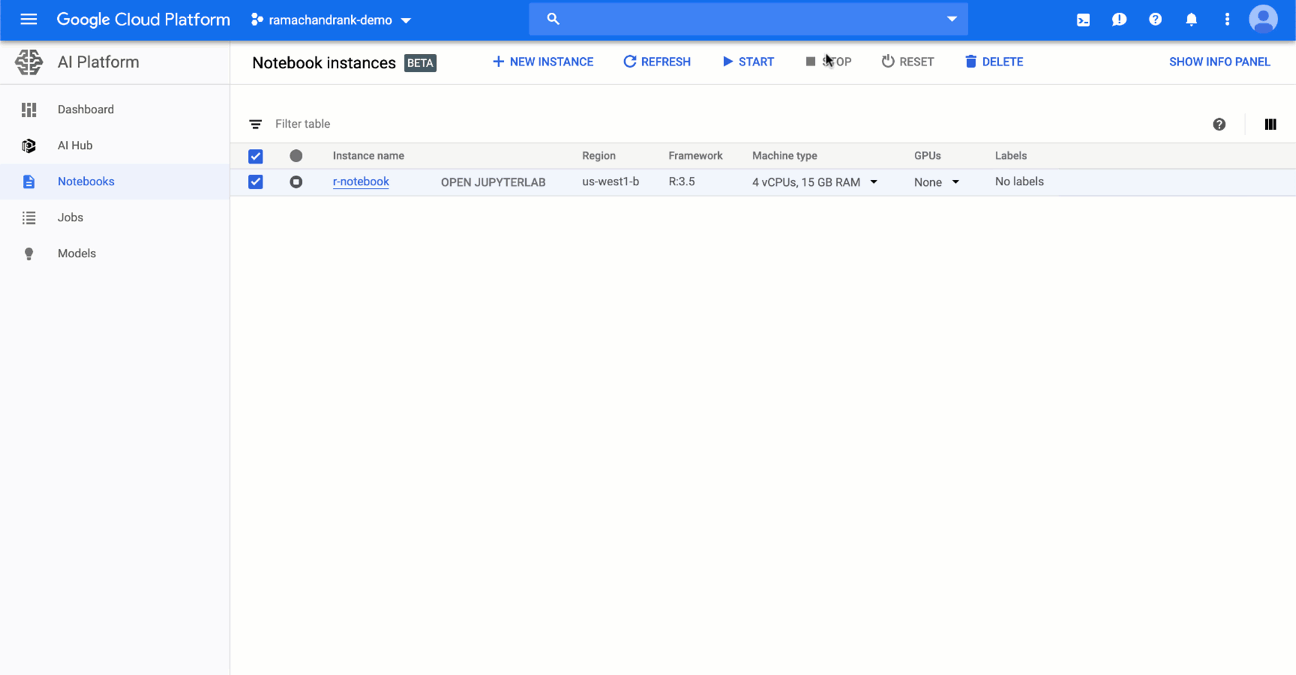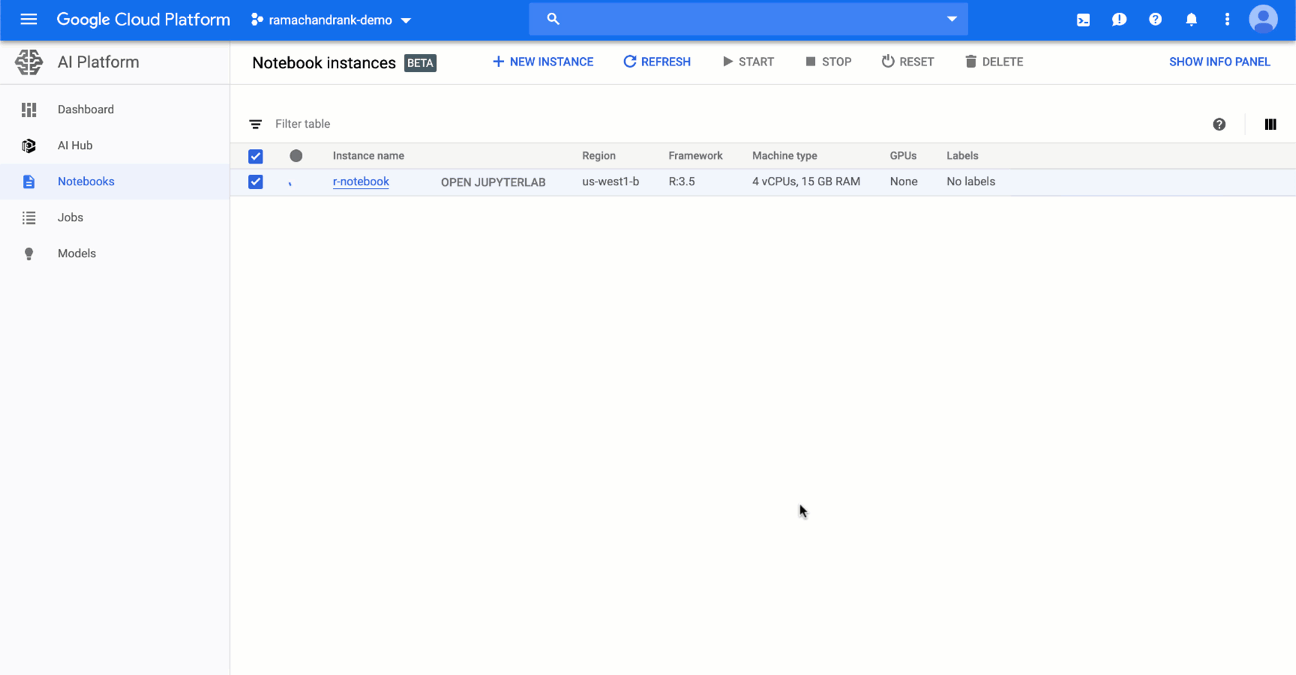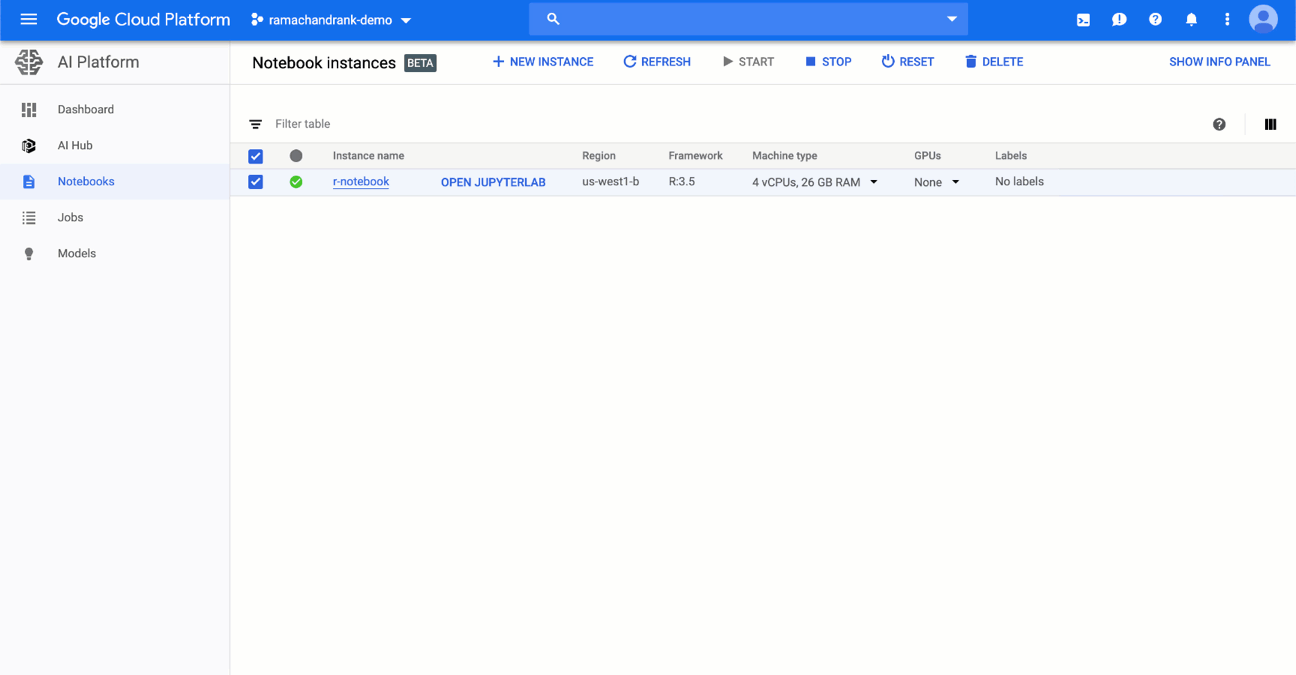
AI Platform Notebooks let you easily scale your Notebook instances up or down. To change the amount of memory and the number of CPUs available to your instance:
1. Stop your instance by clicking on the check box next to the instance and clicking the Stop button.
2. Click on the Machine Type column and change the number of CPUs and amount of RAM available.
3. Review your changes and hit confirm.